
The ROI was centered over the Detection Zone within each well. In the Specify window, “width” and “height” were set at 2mm, and the “oval” box Using the menu command Edit–> Selection–> Specify. Measuring 2mm in diameter (the same diameter as the tip of the stopper) was made Using the binary image, a circular region-of-interest (ROI) Overlapping nuclei were separated by performing a Watershed segmentation Window, the thresholded image was converted to a binary image. Threshold was set for each grayscale image ( Image–>Īdjust–> Threshold). Images were acquired using a 5X objective onĪ Zeiss Axiovert 200 inverted microscope equipped with a CCD camera.Ĭell migration into the Detection Zone was measured byĬounting cell number using ImageJ analysis software (version 1.42l). Afterġ6 hours, cells were fixed with 0.25% glutaraldehyde and cell nuclei were Tissue Culture Treated, Collagen I coated, or Fibronectin coated wells. MDA-MB-231 breast epithelial cells and HT-1080 fibrosarcomaĬells were cultured on an Oris™ Cell Migration Assay – TriCoated plate having Also, check out my article on measuring intracellular fluorescence with ImageJ.Figure 1. ImageJ is a powerful tool and there are hundreds of tutorials even for more advanced work flows. If you’re looking for more ways to automate data processing in the lab, don’t be afraid to search online. To alter the summary measurements, before you analyze the particles in the previous step, select: Analyze > Set Measurements. The summary box provides you with the total number of cells along with other useful measurements. If necessary, select cells and make manual adjustments if non-cell clusters were selected. The ROI manager contains all the cells (or regions of interest) that were selected. 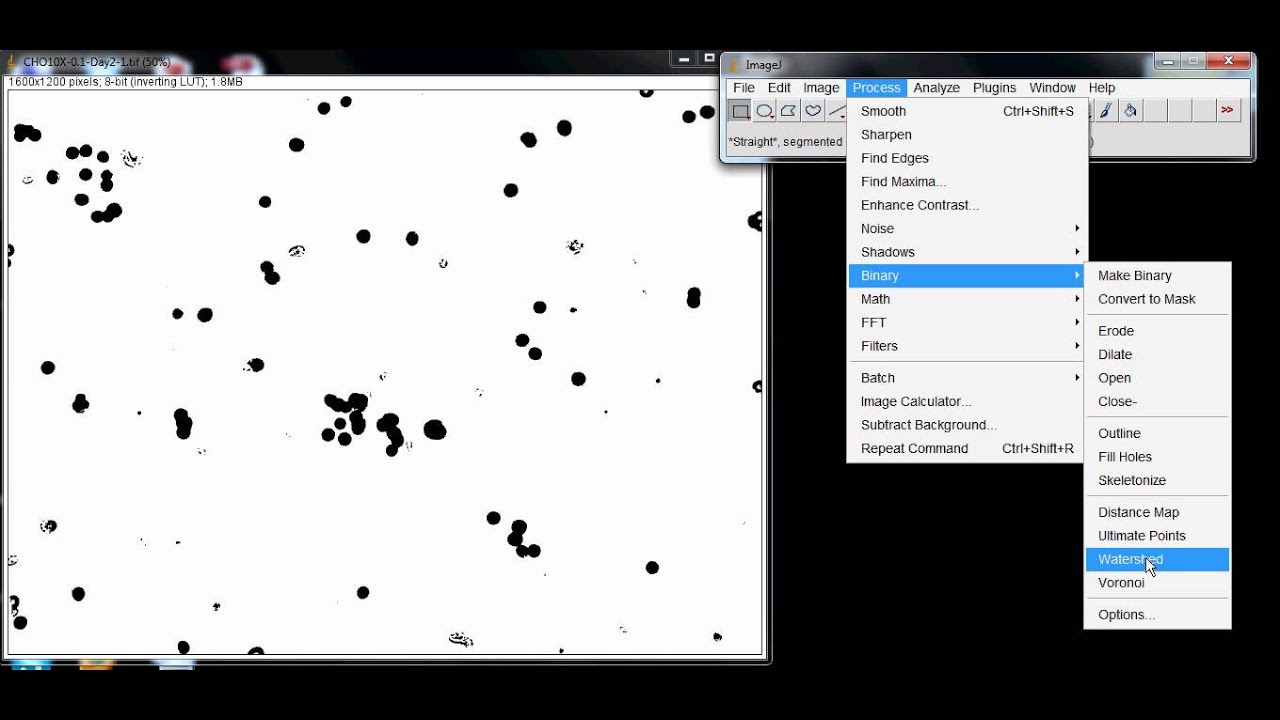
I’d recommend retaining the default setting.Īfter clicking OK in the analyze particles box, the ROI manager and a summary will automatically appear. The second parameter is circularity which asks how circular your cells are. However, if this is the case, set the maximum cell size using the same method just described. Usually you can leave the upper bound as infinity unless multiple cells are packed together. You can then press the key m and a measurement box will popup, providing the area (pixels^2). One way to find a good number is to draw a circle representative of the smallest cell in your image. It’s important to put a lower bound in place to ignore circular substructures within cells. Basically you’re telling the program to ignore any clusters it identifies that are too small to be a cell. To analyze particles, select: Analyze > Analyze Particles. Now that we’ve identified which pixels makeup cells, we’re going to cluster pixels based on circularity using the analyze particles feature. This gave me a binary image in which cell pixels are black while everything else is white. In this example, I removed pixels with an intensity value below 97. Each image is different therefore, you’ll need to play around with the slider to remove as much background as possible without removing cells. The goal is to suppress lower intensity pixels that do not makeup cells therefore set the second slider at 255 and focus on adjusting the lower bound. The first slider determines the lower bound while the second slider determines the upper bound. To adjust threshold, select: Image > Adjust > Threshold. You use this feature to suppress certain pixels in the background by removing intensities below/above a certain threshold. To distinguish cells from background, use the threshold feature. To do this, select: Image > Type > 16-bit Grayscale As you’ll see, we need to convert to grayscale in the first step either way. As long as you can identify the cells against the background you’re in good shape. If you aren’t using colored images, that’s not a problem. Setup for Automatic Cell Countingįirst, load your image by dragging it into the ImageJ toolbox. For those of you unfamiliar with ImageJ, it’s a popular image processing program that runs on Mac, Windows, and Linux.Īssuming you have ImageJ downloaded, let’s begin with a single image of fluorescent cells waiting to be counted. Are you wondering how on earth you’re going to count thousands of cells across a stack of images? Well, I’m going to show you a simple method for automatic cell counting with ImageJ.
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
